5 0 0 0 OA Importance of consensus region of multiple-ligand templates in a virtual screening method
- 著者
- Tatsuya Okuno Koya Kato Shintaro Minami Tomoki P. Terada Masaki Sasai George Chikenji
- 出版者
- 一般社団法人 日本生物物理学会
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.13, pp.149-156, 2016 (Released:2016-07-14)
- 参考文献数
- 31
- 被引用文献数
- 3
We discuss methods and ideas of virtual screening (VS) for drug discovery by examining the performance of VS-APPLE, a recently developed VS method, which extensively utilizes the tendency of single binding pockets to bind diversely different ligands, i.e. promiscuity of binding pockets. In VS-APPLE, multiple ligands bound to a pocket are spatially arranged by maximizing structural overlap of the protein while keeping their relative position and orientation with respect to the pocket surface, which are then combined into a multiple-ligand template for screening test compounds. To greatly reduce the computational cost, comparison of test compound structures are made only with limited regions of the multiple-ligand template. Even when we use the narrow regions with most densely populated atoms for the comparison, VS-APPLE outperforms other conventional VS methods in terms of Area Under the Curve (AUC) measure. This region with densely populated atoms corresponds to the consensus region among multiple ligands. It is typically observed that expansion of the sampled region including more atoms improves screening efficiency. However, for some target proteins, considering only a small consensus region is enough for the effective screening of test compounds. These results suggest that the performance test of VS methods sheds light on the mechanisms of protein-ligand interactions, and elucidation of the protein-ligand interactions should further help improvement of VS methods.
- 著者
- Masaki Sasai George Chikenji Tomoki P. Terada
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.13, pp.281-293, 2016 (Released:2016-11-18)
- 参考文献数
- 73
- 被引用文献数
- 3 5
A simple statistical mechanical model proposed by Wako and Saitô has explained the aspects of protein folding surprisingly well. This model was systematically applied to multiple proteins by Muñoz and Eaton and has since been referred to as the Wako-Saitô-Muñoz-Eaton (WSME) model. The success of the WSME model in explaining the folding of many proteins has verified the hypothesis that the folding is dominated by native interactions, which makes the energy landscape globally biased toward native conformation. Using the WSME and other related models, Saitô emphasized the importance of the hierarchical pathway in protein folding; folding starts with the creation of contiguous segments having a native-like configuration and proceeds as growth and coalescence of these segments. The Φ-values calculated for barnase with the WSME model suggested that segments contributing to the folding nucleus are similar to the structural modules defined by the pattern of native atomic contacts. The WSME model was extended to explain folding of multi-domain proteins having a complex topology, which opened the way to comprehensively understanding the folding process of multi-domain proteins. The WSME model was also extended to describe allosteric transitions, indicating that the allosteric structural movement does not occur as a deterministic sequential change between two conformations but as a stochastic diffusive motion over the dynamically changing energy landscape. Statistical mechanical viewpoint on folding, as highlighted by the WSME model, has been renovated in the context of modern methods and ideas, and will continue to provide insights on equilibrium and dynamical features of proteins.
- 著者
- Sumita Das Tomoki P. Terada Masaki Sasai
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.15, pp.136-150, 2018 (Released:2018-05-26)
- 参考文献数
- 47
- 被引用文献数
- 9
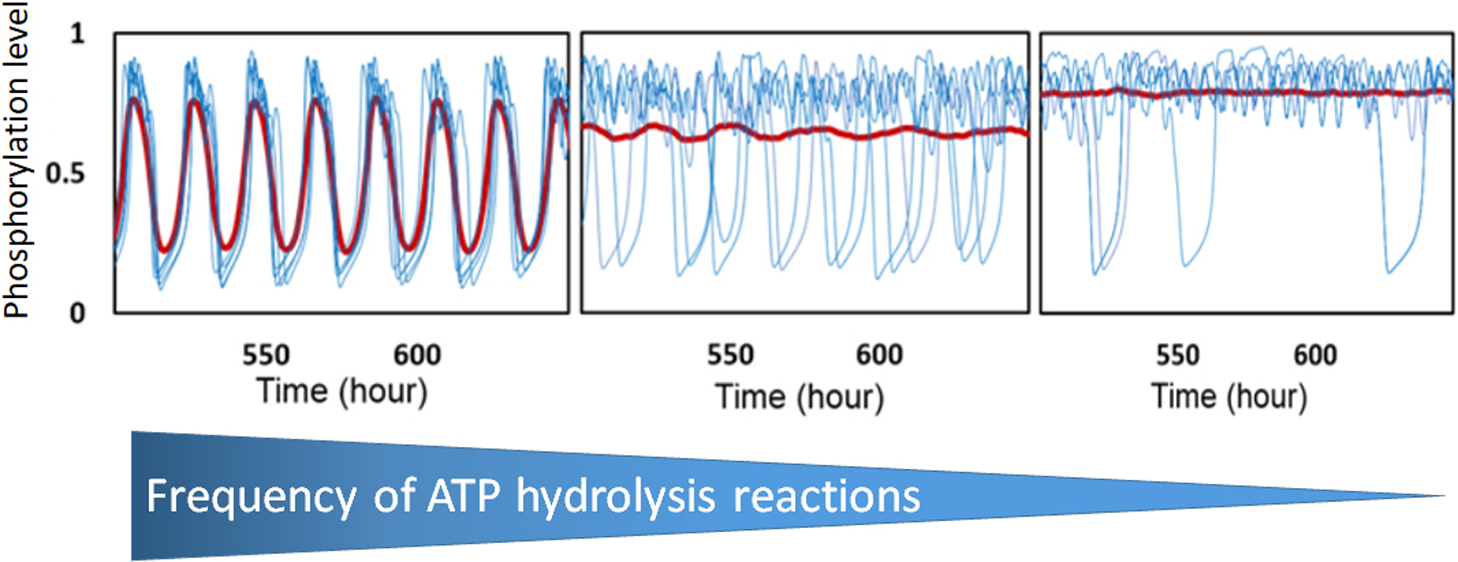
When three cyanobacterial proteins, KaiA, KaiB, and KaiC, are incubated with ATP in vitro, the phosphorylation level of KaiC hexamers shows stable oscillation with approximately 24 h period. In order to understand this KaiABC clockwork, we need to analyze both the macroscopic synchronization of a large number of KaiC hexamers and the microscopic reactions and structural changes in individual KaiC molecules. In the present paper, we explain two coarse-grained theoretical models, the many-molecule (MM) model and the single-molecule (SM) model, to bridge the gap between macroscopic and microscopic understandings. In the simulation results with these models, ATP hydrolysis in the CI domain of KaiC hexamers drives oscillation of individual KaiC hexamers and the ATP hydrolysis is necessary for synchronizing oscillations of a large number of KaiC hexamers. Sensitive temperature dependence of the lifetime of the ADP bound state in the CI domain makes the oscillation period temperature insensitive. ATPase activity is correlated to the frequency of phosphorylation oscillation in the single molecule of KaiC hexamer, which should be the origin of the observed ensemble-level correlation between the ATPase activity and the frequency of phosphorylation oscillation. Thus, the simulation results with the MM and SM models suggest that ATP hydrolysis stochastically occurring in each CI domain of individual KaiC hexamers is a key process for oscillatory behaviors of the ensemble of many KaiC hexamers.