- 著者
- Takeru Kameda Akinori Awazu Yuichi Togashi
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.19, pp.e190027, 2022 (Released:2022-09-08)
- 参考文献数
- 211
- 被引用文献数
- 2
With the recent progress in structural biology and genome biology, structural dynamics of molecular systems that include nucleic acids has attracted attention in the context of gene regulation. The structure–function relationship is an important topic that highlights the importance of the physicochemical properties of nucleotides, as well as that of amino acids in proteins. Simulations are a useful tool for the detailed analysis of molecular dynamics that complement experiments in molecular biology; however, molecular simulation of nucleic acids is less well developed than that of proteins partly due to the physical nature of nucleic acids. In this review, we briefly describe the current status and future directions of the field as a guide to promote collaboration between experimentalists and computational biologists.
- 著者
- Shingo Wakao Noriko Saitoh Akinori Awazu
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.e200020, (Released:2023-04-27)
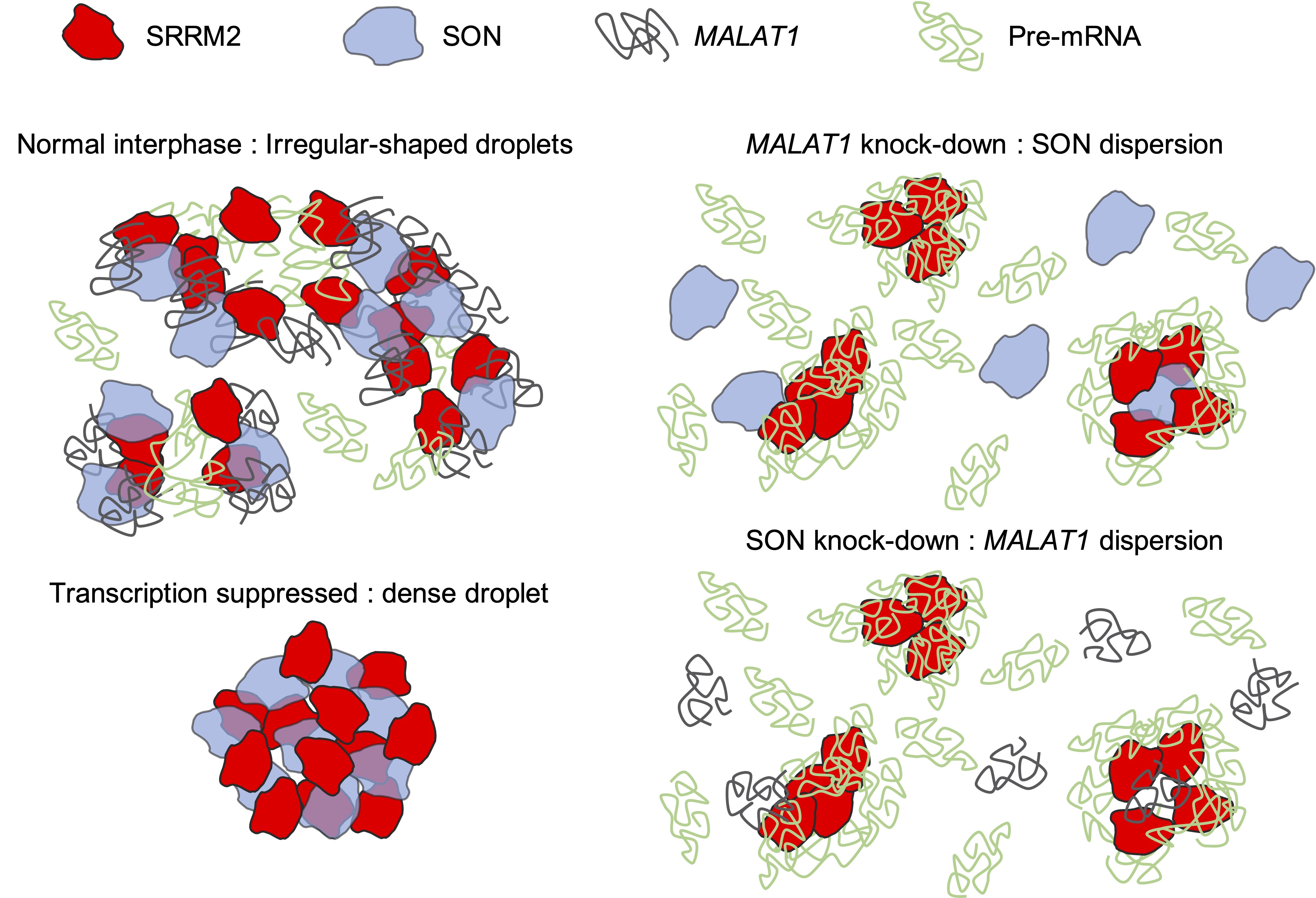
Nuclear speckles are nuclear bodies consisting of populations of small and irregularly shaped droplet-like molecular condensates that contain various splicing factors. Recent experiments have revealed the following structural features of nuclear speckles: (I) Each molecular condensate contains SON and SRRM2 proteins, and MALAT1 non-coding RNA surrounds these condensates; (II) During normal interphase of the cell cycle in multicellular organisms, these condensates are broadly distributed throughout the nucleus. In contrast, when cell transcription is suppressed, the condensates fuse and form strongly condensed spherical droplets; (III) SON is dispersed spatially in MALAT1 knocked-down cells and MALAT1 is dispersed in SON knocked-down cells because of the collapse of the nuclear speckles. However, the detailed interactions among the molecules that are mechanistically responsible for the structural variation remain unknown. In this study, a coarse-grained molecular dynamics model of the nuclear speckle was developed by considering the dynamics of SON, SRRM2, MALAT1, and pre-mRNA as representative components of the condensates. The simulations reproduced the structural changes, which were used to predict the interaction network among the representative components of the condensates.
- 著者
- Tetsushi Komoto Masashi Fujii Akinori Awazu
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.19, pp.e190018, 2022 (Released:2022-06-01)
- 参考文献数
- 48
- 被引用文献数
- 1
X chromosome inactivation center (Xic) pairing occurs during the differentiation of embryonic stem (ES) cells from female mouse embryos, and is related to X chromosome inactivation, the circadian clock, intra-nucleus architecture, and metabolism. However, the mechanisms underlying the identification and approach of X chromosome pairs in the crowded nucleus are unclear. To elucidate the driving force of Xic pairing, we developed a coarse-grained molecular dynamics model of intranuclear chromosomes in ES cells and in cells 2 days after the onset of differentiation (2-day cells) by considering intrachromosomal epigenetic-structural feature-dependent mechanics. The analysis of the experimental data showed that X-chromosomes exhibit the rearrangement of their distributions of open/closed chromatin regions on their surfaces during cell differentiation. By simulating models where the excluded volume effects of closed chromatin regions are stronger than those of open chromatin regions, such rearrangement of open/closed chromatin regions on X-chromosome surfaces promoted the mutual approach of the Xic pair. These findings suggested that local intrachromosomal epigenetic features may contribute to the regulation of cell species-dependent differences in intranuclear architecture.
- 著者
- Shingo Wakao Noriko Saitoh Akinori Awazu
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.20, no.2, pp.e200020, 2023 (Released:2023-05-23)
- 参考文献数
- 33
Nuclear speckles are nuclear bodies consisting of populations of small and irregularly shaped droplet-like molecular condensates that contain various splicing factors. Recent experiments have revealed the following structural features of nuclear speckles: (I) Each molecular condensate contains SON and SRRM2 proteins, and MALAT1 non-coding RNA surrounds these condensates; (II) During normal interphase of the cell cycle in multicellular organisms, these condensates are broadly distributed throughout the nucleus. In contrast, when cell transcription is suppressed, the condensates fuse and form strongly condensed spherical droplets; (III) SON is dispersed spatially in MALAT1 knocked-down cells and MALAT1 is dispersed in SON knocked-down cells because of the collapse of the nuclear speckles. However, the detailed interactions among the molecules that are mechanistically responsible for the structural variation remain unknown. In this study, a coarse-grained molecular dynamics model of the nuclear speckle was developed by considering the dynamics of SON, SRRM2, MALAT1, and pre-mRNA as representative components of the condensates. The simulations reproduced the structural changes, which were used to predict the interaction network among the representative components of the condensates.
2 0 0 0 OA Mathematical model of chromosomal dynamics during DNA double strand break repair in budding yeast
- 著者
- Shinjiro Nakahata Tetsushi Komoto Masashi Fujii Akinori Awazu
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.19, pp.e190012, 2022 (Released:2022-04-20)
- 参考文献数
- 40
- 被引用文献数
- 1
During the repair of double-strand breaks (DSBs) in DNA, active mobilizations for conformational changes in chromosomes have been widely observed in eukaryotes, from yeast to animal and plant cells. DSB-damaged loci in the yeast genome showed increased mobility and relocation to the nuclear periphery. However, the driving forces behind DSB-induced chromatin dynamics remain unclear. In this study, mathematical models of normal and DSB-damaged yeast chromosomes were developed to simulate their structural dynamics. The effects of histone degradation in the whole nucleus and the change in the physical properties of damaged loci due to the binding of SUMOylated repair proteins were considered in the model of DSB-induced chromosomes based on recent experimental results. The simulation results reproduced DSB-induced changes to structural and dynamical features by which the combination of whole nuclear histone degradation and the rigid structure formation of repair protein accumulations on damaged loci were suggested to be primary contributors to the process by which damaged loci are relocated to the nuclear periphery.
- 著者
- Masaki Shirai Takuya Nara Haruko Takahashi Kazuya Takayama Yuan Chen Yudai Hirose Masashi Fujii Akinori Awazu Nobuyoshi Shimoda Yutaka Kikuchi
- 出版者
- The Genetics Society of Japan
- 雑誌
- Genes & Genetic Systems (ISSN:13417568)
- 巻号頁・発行日
- pp.21-00092, (Released:2022-06-18)
- 参考文献数
- 42
CpG methylation of genomic DNA is a well-known repressive epigenetic marker in eukaryotic transcription, and DNA methylation of promoter regions is correlated with gene silencing. In contrast to the promoter regions, the function of DNA methylation during transcription termination remains to be elucidated. A recent study revealed that mouse DNA methyltransferase 3a (Dnmt3a) mainly functions in de novo methylation in the promoter and gene body regions, including transcription termination sites (TTSs), during development. To investigate the relationship between DNA methylation overlapping the TTSs and transcription termination, we performed bioinformatics analysis using six pre-existing Dnmt-/- mouse cell datasets: four types of neurons (three Dnmt3a-/- and one Dnmt1-/- mutants) and two types of embryonic fibroblasts (MEFs) (Dnmt3a-/- and Dnmt3b-/- mutants). Combined analyses using methylome and transcriptome data revealed that read counts downstream of hypomethylated TTSs were increased in three types of neurons (two Dnmt3a-/- and one Dnmt1-/- mutants). Among these, an increase in chimeric transcripts downstream of the TTSs was observed in Dnmt3a-/- mature olfactory sensory neurons and Dnmt3a-/- agouti-related peptide (protein)-producing neurons, thereby indicating that read-through occurs in hypomethylated TTSs at specific gene loci in these two mutants. Conversely, in Dnmt3a-/- MEFs, we detected reductions in read counts downstream of hypomethylated TTSs. These results indicate that the hypomethylation of TTSs can both positively and negatively regulate transcription termination, dependent on Dnmt and cell types. This study is the first to identify the aberrant termination of transcription at specific gene loci with DNA hypomethylated TTSs attributable to Dnmt deficiency.