- 著者
- Tohru Minamino Miki Kinoshita Yusuke V. Morimoto Keiichi Namba
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.e190046, (Released:2022-11-19)
- 被引用文献数
- 3
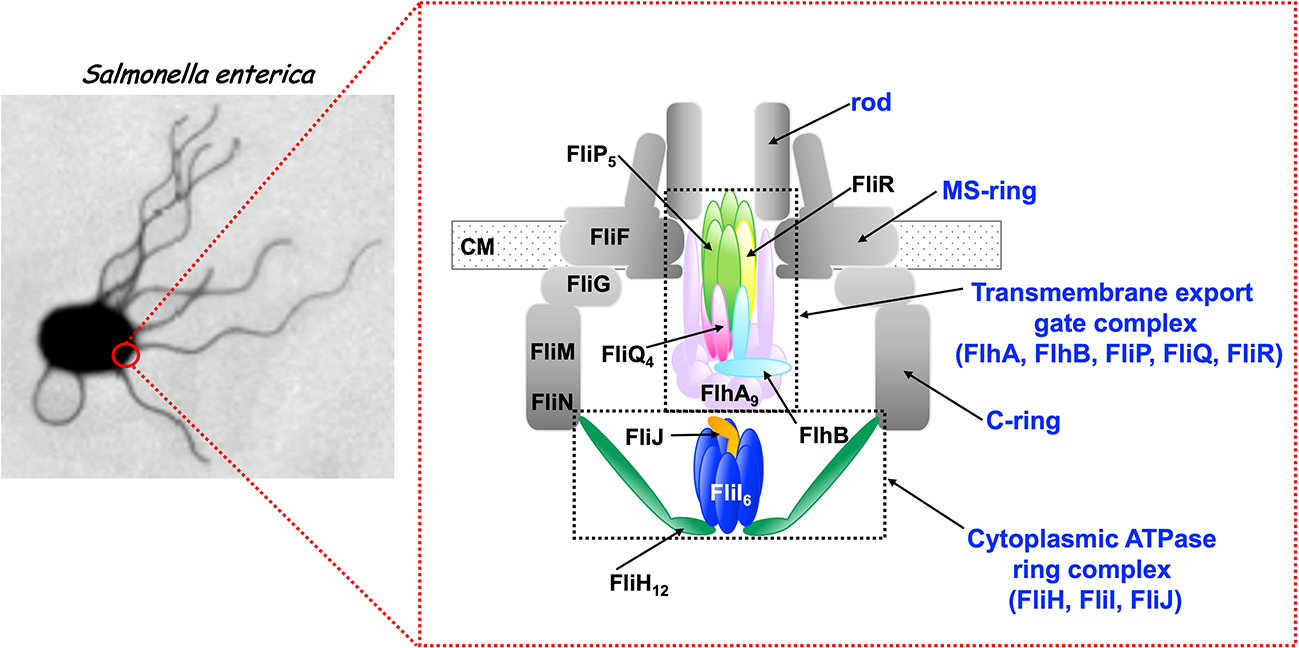
Bacteria employ the flagellar type III secretion system (fT3SS) to construct flagellum, which acts as a supramolecular motility machine. The fT3SS of Salmonella enterica serovar Typhimurium is composed of a transmembrane export gate complex and a cytoplasmic ATPase ring complex. The transmembrane export gate complex is fueled by proton motive force across the cytoplasmic membrane and is divided into four distinct functional parts: a dual-fuel export engine; a polypeptide channel; a membrane voltage sensor; and a docking platform. ATP hydrolysis by the cytoplasmic ATPase complex converts the export gate complex into a highly efficient proton (H+)/ protein antiporter that couples inward-directed H+ flow with outward-directed protein export. When the ATPase ring complex does not work well in a given environment, the export gate complex will remain inactive. However, when the electric potential difference, which is defined as membrane voltage, rises above a certain threshold value, the export gate complex becomes an active H+/protein antiporter to a considerable degree, suggesting that the export gate complex has a voltage-gated activation mechanism. Furthermore, the export gate complex also has a sodium ion (Na+) channel to couple Na+ influx with flagellar protein export. In this article, we review our current understanding of the activation mechanism of the dual-fuel protein export engine of the fT3SS. This review article is an extended version of a Japanese article, Membrane voltage-dependent activation of the transmembrane export gate complex in the bacterial flagellar type III secretion system, published in SEIBUTSU BUTSURI Vol. 62, p165–169 (2022).
- 著者
- Hitomi Komatsu Fumio Hayashi Masahiro Sasa Koji Shikata Shigeru Yamaguchi Keiichi Namba Kenji Oosawa
- 出版者
- 一般社団法人 日本生物物理学会
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.13, pp.13-25, 2016 (Released:2016-01-28)
- 参考文献数
- 81
- 被引用文献数
- 9
FliF is the protein comprising the MS-ring of the bacterial flagellar basal body, which is the base for the assembly of flagellar axial structures. From a fliF mutant that easily releases the rod-hook-filament in viscous environments, more than 400 revertants that recovered their swarming ability in viscous conditions, were isolated. The second-site mutations were determined for approximately 70% of them. There were three regions where the mutations were localized: two in Region I, 112 in Region II, and 71 in Region III including the true reversion. In Region I, second-site mutations were found in FlgC and FlgF of the proximal rod, suggesting that they affect the interaction between the MS-ring and the rod. In Region II, there were 69 and 42 mutations in MotA and MotB, respectively, suggesting that the second-site mutations in MotA and MotB may decrease the rotational speed of the flagellar motor to reduce the probability of releasing the rod under this condition. One exception is a mutation in FlhC that caused a down regulation of the flagellar proteins production but it may directly affect transcription or translation of motA and motB. In Region III, there were 44, 24, and 3 mutations in FliG, FliM, and FliF, respectively. There were no second-site mutations identified in FliN although it is involved in torque generation as a component of the C-ring. Many of the mutations were involved in the motor rotation, and it is suggested that such reduced speeds result in stabilizing the filament attachment to the motor.
- 著者
- Tohru Minamino Miki Kinoshita Yusuke V. Morimoto Keiichi Namba
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.19, pp.e190046, 2022 (Released:2022-12-07)
- 参考文献数
- 83
- 被引用文献数
- 3
Bacteria employ the flagellar type III secretion system (fT3SS) to construct flagellum, which acts as a supramolecular motility machine. The fT3SS of Salmonella enterica serovar Typhimurium is composed of a transmembrane export gate complex and a cytoplasmic ATPase ring complex. The transmembrane export gate complex is fueled by proton motive force across the cytoplasmic membrane and is divided into four distinct functional parts: a dual-fuel export engine; a polypeptide channel; a membrane voltage sensor; and a docking platform. ATP hydrolysis by the cytoplasmic ATPase complex converts the export gate complex into a highly efficient proton (H+)/protein antiporter that couples inward-directed H+ flow with outward-directed protein export. When the ATPase ring complex does not work well in a given environment, the export gate complex will remain inactive. However, when the electric potential difference, which is defined as membrane voltage, rises above a certain threshold value, the export gate complex becomes an active H+/protein antiporter to a considerable degree, suggesting that the export gate complex has a voltage-gated activation mechanism. Furthermore, the export gate complex also has a sodium ion (Na+) channel to couple Na+ influx with flagellar protein export. In this article, we review our current understanding of the activation mechanism of the dual-fuel protein export engine of the fT3SS. This review article is an extended version of a Japanese article, Membrane voltage-dependent activation of the transmembrane export gate complex in the bacterial flagellar type III secretion system, published in SEIBUTSU BUTSURI Vol. 62, p165–169 (2022).
- 著者
- David J. Castillo Shuichi Nakamura Yusuke V. Morimoto Yong-Suk Che Nobunori Kami-ike Seishi Kudo Tohru Minamino Keiichi Namba
- 出版者
- 日本生物物理学会
- 雑誌
- BIOPHYSICS (ISSN:13492942)
- 巻号頁・発行日
- vol.9, pp.173-181, 2013 (Released:2013-12-26)
- 参考文献数
- 46
- 被引用文献数
- 4 34
The bacterial flagellar motor is made of a rotor and stators. In Salmonella it is thought that about a dozen MotA/B complexes are anchored to the peptidoglycan layer around the motor through the C-terminal peptidoglycan-binding domain of MotB to become active stators as well as proton channels. MotB consists of 309 residues, forming a single transmembrane helix (30-50), a stalk (51-100) and a C-terminal peptidoglycan-binding domain (101-309). Although the stalk is dispensable for torque generation by the motor, it is required for efficient motor performance. Residues 51 to 72 prevent premature proton leakage through the proton channel prior to stator assembly into the motor. However, the role of residues 72-100 remains unknown. Here, we analyzed the torque-speed relationship of the MotB(Δ72-100) motor. At a low speed near stall, this mutant motor produced torque at the wild-type level. Unlike the wild-type motor, however, torque dropped off drastically by slight decrease in external load and then showed a slow exponential decay over a wide range of load by its further reduction. Since it is known that the stator is a mechanosensor and that the number of active stators changes in a load-dependent manner, we interpreted this unusual torque-speed relationship as anomaly in load-dependent control of the number of active stators. The results suggest that residues 72-100 of MotB is required for proper load-dependent control of the number of active stators around the rotor.