- 著者
- Toshiki Yagi Akiyuki Toda Muneyoshi Ichikawa Genji Kurisu
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.e200008, (Released:2023-02-08)
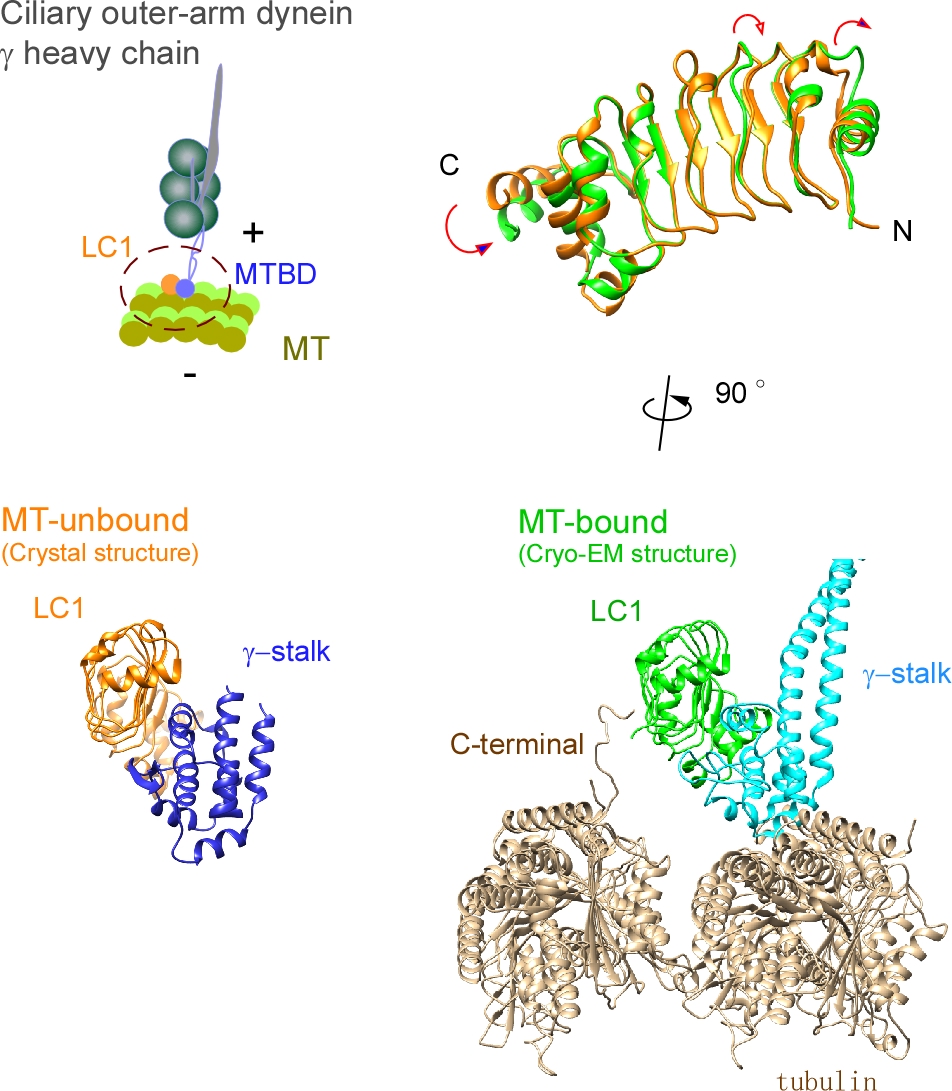
Ciliary bending movements are powered by motor protein axonemal dyneins. They are largely classified into two groups, inner-arm dynein and outer-arm dynein. Outer-arm dynein, which is important for the elevation of ciliary beat frequency, has three heavy chains (α, β, and γ), two intermediate chains, and more than 10 light chains in green algae, Chlamydomonas. Most of intermediate chains and light chains bind to the tail regions of heavy chains. In contrast, the light chain LC1 was found to bind to the ATP-dependent microtubule-binding domain of outer-arm dynein γ-heavy chain. Interestingly, LC1 was also found to interact with microtubules directly, but it reduces the affinity of the microtubule-binding domain of γ-heavy chain for microtubules, suggesting the possibility that LC1 may control ciliary movement by regulating the affinity of outer-arm dyneins for microtubules. This hypothesis is supported by the LC1 mutant studies in Chlamydomonas and Planaria showing that ciliary movements in LC1 mutants were disordered with low coordination of beating and low beat frequency. To understand the molecular mechanism of the regulation of outer-arm dynein motor activity by LC1, X-ray crystallography and cryo-electron microscopy have been used to determine the structure of the light chain bound to the microtubule-binding domain of γ-heavy chain. In this review article, we show the recent progress of structural studies of LC1, and suggest the regulatory role of LC1 in the motor activity of outer-arm dyneins. This review article is an extended version of the Japanese article, The Complex of Outer-arm Dynein Light Chain-1 and the Microtubule-binding Domain of the Heavy Chain Shows How Axonemal Dynein Tunes Ciliary Beating, published in SEIBUTSU BUTSURI Vol. 61, p.20-22 (2021).
- 著者
- Yuji Furutani Chii-Shen Yang
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.e201005, (Released:2023-01-11)
- 著者
- Kumiko Hayashi Jakia Jannat Keya
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.e190030, (Released:2022-08-27)
- 被引用文献数
- 1
- 著者
- Madoka Suzuki Kotaro Oyama
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.e190029, (Released:2022-08-27)
- 被引用文献数
- 1
- 著者
- Damien Simon Atsushi Mukaiyama Yoshihiko Furuike Shuji Akiyama
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.e190008, (Released:2022-03-30)
- 被引用文献数
- 5
- 著者
- Keisuke Inoue Shoji Takada Tsuyoshi Terakawa
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.19, pp.e190015, 2022 (Released:2022-05-11)
- 参考文献数
- 43
- 被引用文献数
- 2
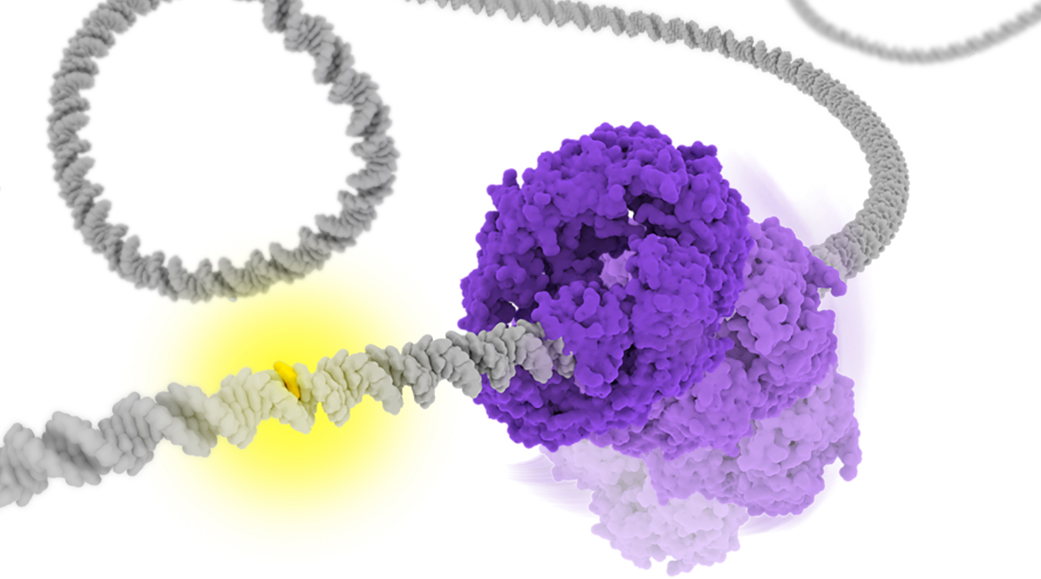
DNA mismatches are frequently generated by various intrinsic and extrinsic factors including DNA replication errors, oxygen species, ultraviolet, and ionizing radiation. These mismatches should be corrected by the mismatches repair (MMR) pathway to maintain genome integrity. In the Escherichia coli (E. coli) MMR pathway, MutS searches and recognizes a base-pair mismatch from millions of base-pairs. Once recognized, ADP bound to MutS is exchanged with ATP, which induces a conformational change in MutS. Previous single-molecule fluorescence microscopy studies have suggested that ADP-bound MutS temporarily slides along double-stranded DNA in a rotation-coupled manner to search a base-pair mismatch and so does ATP-bound MutS in a rotation-uncoupled manner. However, the detailed structural dynamics of the sliding remains unclear. In this study, we performed coarse-grained molecular dynamics simulations of the E. coli MutS bound on DNA in three different conformations: ADP-bound (), ATP-bound open clamp (), and ATP-bound closed clamp () conformations. In the simulations, we observed conformation-dependent diffusion of MutS along DNA. and diffused along DNA in a rotation-coupled manner with rare and frequent groove-crossing events, respectively. In the groove-crossing events, MutS overcame an edge of a groove and temporarily diffused in a rotation-uncoupled manner. It was also indicated that mismatch searches by is inefficient in terms of mismatch checking even though it diffuses along DNA and reaches unchecked regions more rapidly than .
2 0 0 0 OA Beyond multi-disciplinary and cross-scale analyses of the cyanobacterial circadian clock system
- 著者
- Shuji Akiyama Hironari Kamikubo
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.18, pp.267-268, 2021 (Released:2021-11-27)
- 参考文献数
- 10
2 0 0 0 OA Limitations of the ABEGO-based backbone design: ambiguity between αα-corner and αα-hairpin
- 著者
- Koya Sakuma
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.bppb-v18.017, (Released:2021-05-28)
- 著者
- Yutaka Maruyama Hiroshi Takano Ayori Mitsutake
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.16, pp.407-429, 2019 (Released:2019-11-29)
- 参考文献数
- 182
- 被引用文献数
- 6
Molecular dynamics simulation is a fruitful tool for investigating the structural stability, dynamics, and functions of biopolymers at an atomic level. In recent years, simulations can be performed on time scales of the order of milliseconds using specialpurpose systems. Since the most stable structure, as well as meta-stable structures and intermediate structures, is included in trajectories in long simulations, it is necessary to develop analysis methods for extracting them from trajectories of simulations. For these structures, methods for evaluating the stabilities, including the solvent effect, are also needed. We have developed relaxation mode analysis to investigate dynamics and kinetics of simulations based on statistical mechanics. We have also applied the three-dimensional reference interaction site model theory to investigate stabilities with solvent effects. In this paper, we review the results for designing amino-acid substitution of the 10-residue peptide, chignolin, to stabilize the misfolded structure using these developed analysis methods.
- 著者
- Teppei Sugimoto Kota Katayama Hideki Kandori
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.bppb-v18.012, (Released:2021-04-16)
- 被引用文献数
- 6
- 著者
- Akira R. Kinjo
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.12, pp.117-119, 2015 (Released:2015-12-11)
- 参考文献数
- 16
- 被引用文献数
- 1 1
The direct-coupling analysis is a powerful method for protein contact prediction, and enables us to extract “direct” correlations between distant sites that are latent in “indirect” correlations observed in a protein multiple-sequence alignment. I show that the direct correlation can be obtained by using a formulation analogous to the Ornstein-Zernike integral equation in liquid theory. This formulation intuitively illustrates how the indirect or apparent correlation arises from an infinite series of direct correlations, and provides interesting insights into protein structure prediction.
- 著者
- Yujiro Nagasaka Shoko Hososhima Naoko Kubo Takashi Nagata Hideki Kandori Keiichi Inoue Hiromu Yawo
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- pp.BSJ-2020007, (Released:2020-06-09)
- 被引用文献数
- 5
2 0 0 0 OA Consistency principle for protein design
- 著者
- Rie Koga Nobuyasu Koga
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.16, pp.304-309, 2019 (Released:2019-11-29)
- 参考文献数
- 38
- 被引用文献数
- 7
Protein design holds promise for applications such as the control of cells, therapeutics, new enzymes and protein-based materials. Recently, there has been progress in rational design of protein molecules, and a lot of attempts have been made to create proteins with functions of our interests. The key to the progress is the development of methods for controlling desired protein tertiary structures with atomic-level accuracy. A theory for protein folding, the consistency principle, proposed by Nobuhiro Go in 1983, was a compass for the development. Anfinsen hypothesized that proteins fold into the free energy minimum structures, but Go further considered that local and non-local interactions in the free energy minimum structures are consistent with each other. Guided by the principle, we proposed a set of rules for designing ideal protein structures stabilized by consistent local and non-local interactions. The rules made possible designs of amino acid sequences with funnel-shaped energy landscapes toward our desired target structures. So far, various protein structures have been created using the rules, which demonstrates significance of our rules as intended. In this review, we briefly describe how the consistency principle impacts on our efforts for developing the design technology.
- 著者
- Yasumasa Joti Akio Kitao
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.16, pp.240-247, 2019 (Released:2019-11-29)
- 参考文献数
- 28
- 被引用文献数
- 2
Terahertz time-domain spectra (THz-TDS) were investigated using the results of molecular dynamics (MD) simulations of Staphylococcal nuclease at two hydration states in the temperature range between 100 and 300 K. The temperature dependence of THz-TDS was found to differ significantly from that of the incoherent neutron scattering spectra (INSS) calculated from the same MD simulation results. We further examined contributions of the mutual and auto-correlations of the atomic fluctuations to THz-TDS and found that the negative value of the former contribution nearly canceled out the positive value of the latter, resulting in a monotonic increase of the reduced absorption cross section. Because of this cancellation, no distinct broad peak was observed in the absorption lineshape function of THz-TDS, whereas the protein boson peak was observed in INSS. The contribution of water molecules to THz-TDS was extremely large for the hydrated protein at temperatures above 200 K, in which large-amplitude motions of water were excited. The combination of THz-TDS, INSS and MD simulations has the potential to extract function-relevant protein dynamics occurring on the picosecond to nanosecond timescale.
- 著者
- Kazuhiro Takemura Akio Kitao
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.16, pp.295-303, 2019 (Released:2019-11-29)
- 参考文献数
- 36
- 被引用文献数
- 4
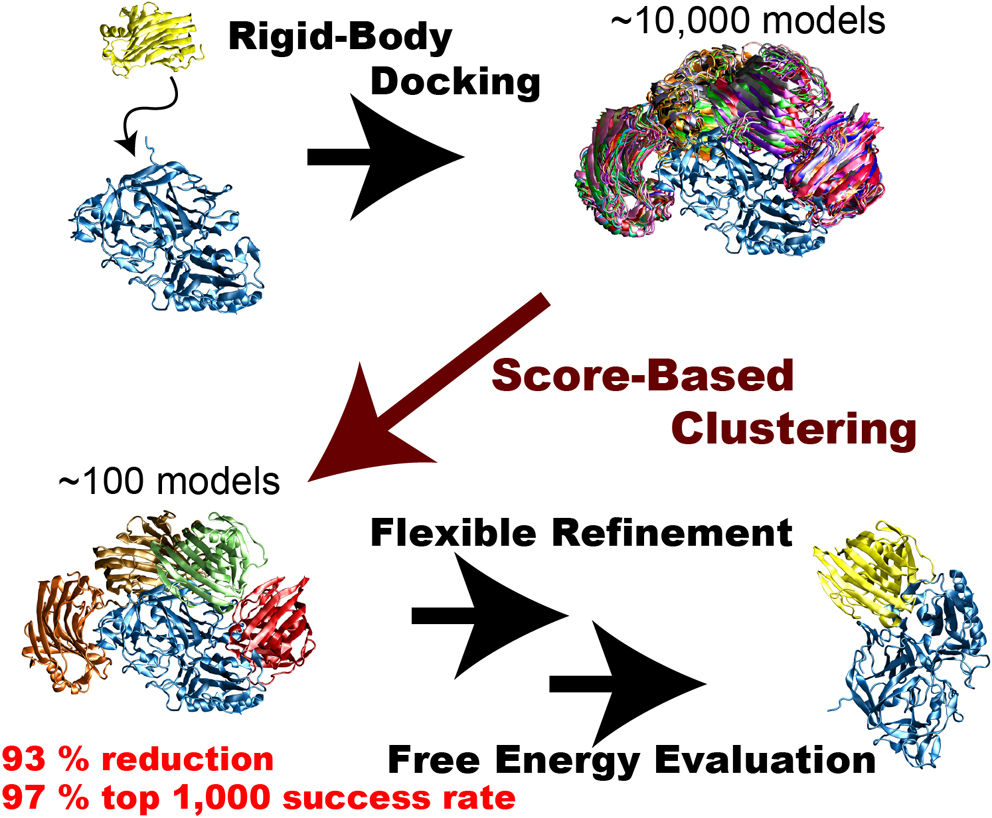
Rigid-body protein-protein docking is very efficient in generating tens of thousands of docked complex models (decoys) in a very short time without considering structure change upon binding, but typical docking scoring functions are not necessarily sufficiently accurate to narrow these decoys down to a small number of plausible candidates. Flexible refinements and sophisticated evaluation of the decoys are thus required to achieve more accurate prediction. Since this process is time-consuming, an efficient screening method to reduce the number of decoys is necessary immediately following rigid-body dockings. We attempted to develop an efficient screening method by clustering decoys generated by the rigid-body docking ZDOCK. We introduced the three metrics ligand-root-mean-square deviation (L-RMSD), interface-ligand-RMSD (iL-RMSD), and the fraction of common contacts (FCC), and examined various ranges of cut-offs for clusters to determine the best set of clustering parameters. Although the employed clustering algorithm is simple, it successfully reduced the number of decoys. Using iL-RMSD with a cut-off radius of 8 Å, the number of decoys that contain at least one near-native model with 90% probability decreased from 4,808 to 320, a 93% reduction in the original number of decoys. Using FCC for the clustering step, the top 1,000 success rates, defined as the probability that the top 1,000 models contain at least one near-native structure, reached 97%. We conclude that the proposed method is very efficient in selecting a small number of decoys that include near-native decoys.
- 著者
- Hiroyuki Terashima Akihiro Kawamoto Yusuke V. Morimoto Katsumi Imada Tohru Minamino
- 出版者
- The Biophysical Society of Japan
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.14, pp.191-198, 2017 (Released:2017-12-19)
- 参考文献数
- 57
- 被引用文献数
- 1 43
The bacterial flagellum is a supramolecular motility machine consisting of the basal body as a rotary motor, the hook as a universal joint, and the filament as a helical propeller. Intact structures of the bacterial flagella have been observed for different bacterial species by electron cryotomography and subtomogram averaging. The core structures of the basal body consisting of the C ring, the MS ring, the rod and the protein export apparatus, and their organization are well conserved, but novel and divergent structures have also been visualized to surround the conserved structure of the basal body. This suggests that the flagellar motors have adapted to function in various environments where bacteria live and survive. In this review, we will summarize our current findings on the divergent structures of the bacterial flagellar motor.
2 0 0 0 OA What parameters characterize “life”?
- 著者
- Shigeki Mitaku Ryusuke Sawada
- 出版者
- 一般社団法人 日本生物物理学会
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.13, pp.305-310, 2016 (Released:2016-11-18)
- 参考文献数
- 7
- 被引用文献数
- 1 5
“Life” is a particular state of matter, and matter is composed of various molecules. The state corresponding to “life” is ultimately determined by the genome sequence, and this sequence determines the conditions necessary for survival of the organism. In order to elucidate one parameter characterizing the state of “life”, we analyzed the amino acid sequences encoded in the total genomes of 557 prokaryotes and 40 eukaryotes using a membrane protein prediction online tool called SOSUI. SOSUI uses only the physical parameters of the encoded amino acid sequences to make its predictions. The ratio of membrane proteins in a genome predicted by the SOSUI online tool was around 23% for all genomes, indicating that this parameter is controlled by some mechanism in cells. In order to identify the property of genome DNA sequences that is the possible cause of the constant ratio of membrane proteins, we analyzed the nucleotide compositions at codon positions and observed the existence of systematic biases distinct from those expected based on random distribution. We hypothesize that the constant ratio of membrane proteins is the result of random mutations restricted by the systematic biases inherent to nucleotide codon composition. A new approach to the biological sciences based on the holistic analysis of whole genomes is discussed in order to elucidate the principles underlying “life” at the biological system level.
- 著者
- Takako Sakano Md. Iqbal Mahamood Takefumi Yamashita Hideaki Fujitani
- 出版者
- 一般社団法人 日本生物物理学会
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.13, pp.181-194, 2016 (Released:2016-07-14)
- 参考文献数
- 39
- 被引用文献数
- 46
The accurate prediction of a ligand–protein complex structure is important for computer-assisted drug development. Although many docking methods have been developed over the last three decades, the success of binding structure prediction remains greatly limited. The purpose of this study was to demonstrate the usefulness of molecular dynamics (MD) simulation in assessing a docking pose predicted using a docking program. If the predicted pose is not unstable in an aqueous environment, MD simulation equilibrates the system and removes the ligand from the predicted position. Here we investigated two proteins that are important potential therapeutic targets: β2 adrenergic receptor (β2AR) and PR-Set7. While β2AR is rigid and its ligands are very similar to the template ligand (carazolol), PR-Set7 is very flexible and its ligands vary greatly from the template ligand (histone H4 tail peptide). On an empirical basis, we usually expect that the docking prediction is accurate when the protein is rigid and its ligands are similar to the template ligand. The MD analyses in this study clearly suggested such a tendency. Furthermore, we discuss the possibility that the MD simulation can predict the binding pose of a ligand.
- 著者
- Ichiro Yamato Yoshimi Kakinuma Takeshi Murata
- 出版者
- 一般社団法人 日本生物物理学会
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.13, pp.37-44, 2016 (Released:2016-02-27)
- 参考文献数
- 39
- 被引用文献数
- 8
Among the many types of bioenergy-transducing machineries, F- and V-ATPases are unique bio- and nano-molecular rotary motors. The rotational catalysis of F1-ATPase has been investigated in detail, and molecular mechanisms have been proposed based on the crystal structures of the complex and on extensive single-molecule rotational observations. Recently, we obtained crystal structures of bacterial V1-ATPase (A3B3 and A3B3DF complexes) in the presence and absence of nucleotides. Based on these new structures, we present a novel model for the rotational catalysis mechanism of V1-ATPase, which is different from that of F1-ATPases.
- 著者
- Hitomi Komatsu Fumio Hayashi Masahiro Sasa Koji Shikata Shigeru Yamaguchi Keiichi Namba Kenji Oosawa
- 出版者
- 一般社団法人 日本生物物理学会
- 雑誌
- Biophysics and Physicobiology (ISSN:21894779)
- 巻号頁・発行日
- vol.13, pp.13-25, 2016 (Released:2016-01-28)
- 参考文献数
- 81
- 被引用文献数
- 9
FliF is the protein comprising the MS-ring of the bacterial flagellar basal body, which is the base for the assembly of flagellar axial structures. From a fliF mutant that easily releases the rod-hook-filament in viscous environments, more than 400 revertants that recovered their swarming ability in viscous conditions, were isolated. The second-site mutations were determined for approximately 70% of them. There were three regions where the mutations were localized: two in Region I, 112 in Region II, and 71 in Region III including the true reversion. In Region I, second-site mutations were found in FlgC and FlgF of the proximal rod, suggesting that they affect the interaction between the MS-ring and the rod. In Region II, there were 69 and 42 mutations in MotA and MotB, respectively, suggesting that the second-site mutations in MotA and MotB may decrease the rotational speed of the flagellar motor to reduce the probability of releasing the rod under this condition. One exception is a mutation in FlhC that caused a down regulation of the flagellar proteins production but it may directly affect transcription or translation of motA and motB. In Region III, there were 44, 24, and 3 mutations in FliG, FliM, and FliF, respectively. There were no second-site mutations identified in FliN although it is involved in torque generation as a component of the C-ring. Many of the mutations were involved in the motor rotation, and it is suggested that such reduced speeds result in stabilizing the filament attachment to the motor.